Successfully Added to Cart
Customers also bought...
-
 DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the...
DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the... -
 DNA/RNA Shield SafeCollect Swab Collection Kit (1 ml fill) (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA...
DNA/RNA Shield SafeCollect Swab Collection Kit (1 ml fill) (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA...
Highlights
- Quick & Easy: Simple 20 minute procedure.
- Highest Yield: Recover 3x more DNA.
- Ultra-Pure: Ready for qPCR, Next-Gen Sequencing, arrays, etc.
Original Manufacturer
100% satisfaction guaranteed, read Our Promise
Innovated in California, Made in the USA
Highlights
- Quick & Easy: Simple 20 minute procedure.
- Highest Yield: Recover 3x more DNA.
- Ultra-Pure: Ready for qPCR, Next-Gen Sequencing, arrays, etc.
Original Manufacturer
100% satisfaction guaranteed, read Our Promise
Innovated in California, Made in the USA
| Cat # | Name | Size | Price | |
|---|---|---|---|---|
| C1001-50 | Collection Tubes | 50 Pack | $18.00 | |
| D3004-2-50 | g-DNA Wash Buffer | 50 ml | $22.00 | |
| D3004-4-1 | DNA Elution Buffer | 1 ml | $13.00 | |
| D3004-4-10 | DNA Elution Buffer | 10 ml | $17.00 | |
| D3004-2-200 | g-DNA Wash Buffer | 200 ml | $65.00 | |
| D3004-4-50 | DNA Elution Buffer | 50 ml | $39.00 | |
| D3004-5-50 | DNA Pre-Wash Buffer | 50 ml | $31.00 | |
| D3004-5-30 | DNA Pre-Wash Buffer | 30 ml | $25.00 | |
| D3001-2-20 | Proteinase K w/ Storage Buffer Set | 20 mg | $53.00 | |
| D3001-2-5 | Proteinase K w/ Storage Buffer Set | 5 mg | $25.00 | |
| D4068-1-12 | BioFluid & Cell Buffer (Red) | 12 ml | $29.00 | |
| D4068-1-45 | BioFluid & Cell Buffer (Red) | 45 ml | $71.00 | |
| D4068-2-22 | Solid Tissue Buffer (Blue) | 22 ml | $46.00 | |
| D4068-2-6 | Solid Tissue Buffer (Blue) | 6 ml | $14.00 | |
| D4068-3-25 | Genomic Binding Buffer | 25 ml | $34.00 | |
| D4068-3-85 | Genomic Binding Buffer | 85 ml | $114.00 | |
| C1103-50 | Zymo-Spin IC-XM Columns | 50 Pack | $76.00 | |
| C1104-50 | Zymo-Spin IIC-XLR Columns | 50 Pack | $79.00 | |
Description
Performance
Technical Specifications
| Elution Volume | ≥ 35 µl |
|---|---|
| Equipment | Water bath or heat block (55°C), microcentrifuge, and vortex. |
| Purity | High quality DNA is ready for all sensitive downstream applications such as PCR, endonuclease digestion, Southern blotting, genotyping, Next-Gen sequencing, bisulfite conversion, etc. (A260/A230≥ 2.0). |
| Size Range | Capable of recovering genomic and mitochondrial DNA sized fragments > 50 kb. If present, parasitic, microbial, and viral DNA will also be recovered. |
| Workflow | Utilize a Proteinase K Digestion and Zymo-Spin Column for effective recovery of DNA. |
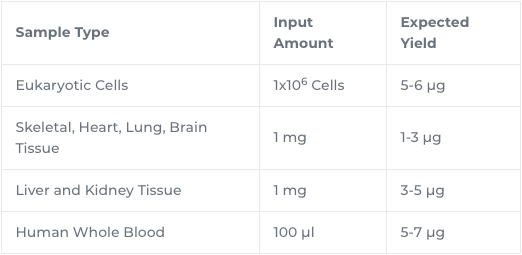
| Yield | The DNA binding capacity of the column is 25 µg. Typically, mammalian tissues yield: 1-3 µg DNA per mg skeletal, heart, lung, and brain tissues and 3-5 µg DNA per mg liver and kidney. Human whole blood will yield 3-7 µg DNA per 100 µl blood sampled.
 |
Resources
Documents
FAQ
The BioFluid & Cell Buffer (Red) is designed to allow for rapid Proteinase K digestion with easy-to-lyse samples (e.g. mammalian cells and biological fluids). The Solid Tissue buffer (Blue) can be used for any sample type and requires a longer PK digestion time compared to using the BioFluid & Cell Buffer (Red).
The Quick-DNA is optimized for cells, soft tissues, and homogenized/digested samples using a single lysis/binding buffer. The Quick-DNA Plus kits contain an optimized Proteinase K for processing a wider variety of sample inputs, such as cells, blood, tissues, etc. The upgraded Quick-DNA Plus recovers more DNA with higher purity compared to the Quick-DNA Kits.
Yes, samples can be digested overnight. Make sure to follow the appendix (page 8 appendix B) for processing liquids samples in DNA/RNA Shield and incubate at room temperature.
White blood cells, which are the major source of genomic DNA in blood, easily and quickly settles. Mix the blood sample well prior to aliquoting for purification.
E.coli cells are easy-to-lyse and can be processed directly with the Biological Fluids & Cells protocol. For an all-inclusive kit with any type of microbes (including tough-to-lyse), use any of Zymo Research’s Environmental Kits (e.g. Quick-DNA Fungal/Bacterial, Quick-DNA Fecal/Soil, ZymoBIOMICS DNA, etc.).
No, the kit is designed for direct use with biological samples. For clean-up of previously isolated DNA, please use the Genomic DNA Clean & Concentrator or the DNA Clean & Concentrator kits.
Beta-mercaptoethanol is a reducing agent that helps break down proteins and improves DNA recovery and purity. Addition of beta-mercaptoethanol is recommended to enhance sample lysis, but can be substituted with dithiothreitol (DTT, final concentration of 10 mM) or omitted.
Yes, add 4 volumes of Genomic Lysis Buffer to 1 volume of crude lysate, homogenized, or digested sample (see Cell Suspensions and Proteinase K Digested Samples) and proceed with the remainder of the protocol.
Reviews
"This kit is easy to use and provides a high quality yield."
M.B.
Syracuse University
"I was very impressed with the purity of the DNA."
C.V.
UCLA
"Improved yield than Bioline and Qiagen Kits."
F.C.
Manchester Metropolitan University
Citations
Need help? Contact Us




