Successfully Added to Cart
Customers also bought...
-
 DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the...
DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the... -
 DNA/RNA Shield SafeCollect Swab Collection Kit (1 ml fill) (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA...
DNA/RNA Shield SafeCollect Swab Collection Kit (1 ml fill) (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA...
Highlights
- Effectively preserves DNA and RNA in urine at ambient temperatures
- Facilitates pelleting of both cellular and cell-free nucleic acids from large volume urine samples
- Inhibits microbial growth during long-term (cold-free) storage of urine samples
Original Manufacturer
100% satisfaction guaranteed, read Our Promise
Innovated in California, Made in the USA
Highlights
- Effectively preserves DNA and RNA in urine at ambient temperatures
- Facilitates pelleting of both cellular and cell-free nucleic acids from large volume urine samples
- Inhibits microbial growth during long-term (cold-free) storage of urine samples
Original Manufacturer
100% satisfaction guaranteed, read Our Promise
Innovated in California, Made in the USA
Description
Resources
Documents
FAQ
Urine samples will be stabilized for a minimum of 30 days at ambient temperature.
Yes, UCB will preserve DNA and RNA integrity.
Mix urine thoroughly and centrifuge at 3000 x g for 15 minutes to form a urine pellet. Purify urine nucleic acid directly from urine pellet using Quick-DNA Urine kit(D3061) or Quick-RNA Miniprep Plus kit (R1057, R1058).
Citations
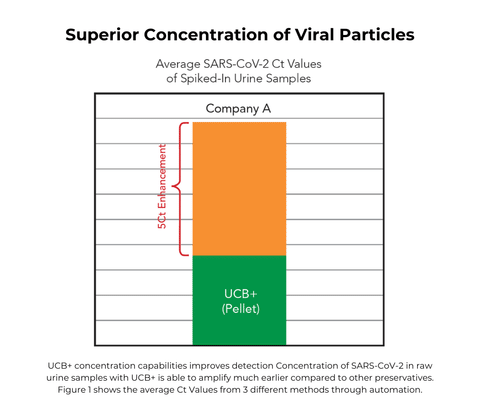
Researchers studying WNV and ZIKV in urine face profound viral RNA loss and major difficulties in viral RNA isolation exhibiting a need for optimal preservation for long-term storage. UCB was used to minimize degradation of WNV RNA in urine during storage. As well as extend the period of detection and detect ZIKV at later time points after infection.
Gorchakov R, et. al. (2019). Optimizing PCR Detection of Zika Virus from Various Body Fluids. Am J Trop Med Hyg. 2019 Feb;100(2):427-433.In an effort to expand the scope of cervical cancer diagnostic procedures, researchers from the Pathology Department of the Cancer Center of Amsterdam compared the hrHPV and DNA methylation results collected from paired urine and cervical scrapes. To ensure that the urine samples they tested were producing the most accurate data possible, Zymo Research Quick DNA Urine Kit with Urine Conditioning Buffer was used with great success.
Snoek BC, et. al (2019). Cervical cancer detection by DNA methylation analysis in urine. Scientific Reports volume 9, Article number: 3088 (2019).Researchers from the Cancer and Genomic Studies Department of the University of Birmingham College of Medical and Dental Sciences conducted a proof-of-principle study investigating the relative quantities of DNA molecules detected by ddPCR exhibiting wild-type and mutant TERT sequences. With little margin for error, the researchers knew the only way to obtain reliable data was to use the Zymo Research Quick-DNA Urine Kit for sample preservation and DNA purification.
Tommasi S, et. al. (2017). Toward Personalized Liquid Biopsies for Urothelial Carcinoma: Characterization of ddPCR and Urinary cfDNA for the Detection of the TERT228 G>A/T Mutation. Urothelial Carcinoma pp 29-41.Read More
Need help? Contact Us