Successfully Added to Cart
Customers also bought...
-
 DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the...
DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the... -
 DNA/RNA Shield SafeCollect Swab Collection Kit (1 ml fill) (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA...
DNA/RNA Shield SafeCollect Swab Collection Kit (1 ml fill) (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA...
Highlights
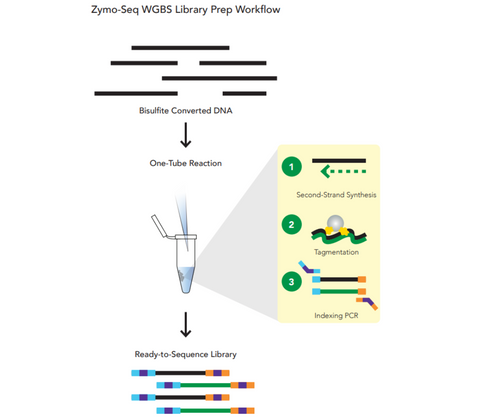
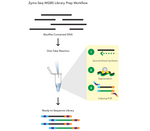
- Whole Genome Bisulfite (WGBS) library preparation in less than 4 hours and in one tube.
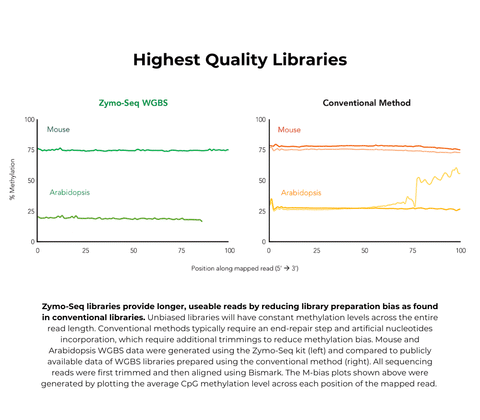
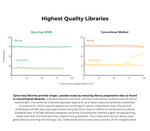
- Reduced library preparation bias for accurate methylation calling
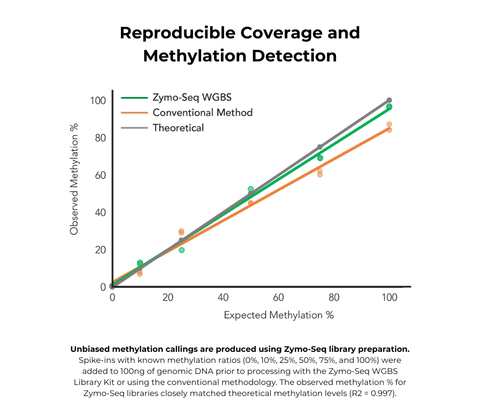
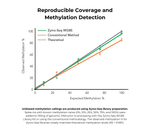
- Reproducible genome coverage from any species (>90% of CpG sites detected in mammalian samples)
Original Manufacturer
Satisfaction 100% guaranteed, read Our Promise
Innovated in California, Made in the USA
Highlights
- Whole Genome Bisulfite (WGBS) library preparation in less than 4 hours and in one tube.
- Reduced library preparation bias for accurate methylation calling
- Reproducible genome coverage from any species (>90% of CpG sites detected in mammalian samples)
Original Manufacturer
Satisfaction 100% guaranteed, read Our Promise
Innovated in California, Made in the USA
| Cat # | Name | Size | Price | |
|---|---|---|---|---|
| D5016 | E. coli Non-Methylated Genomic DNA | 5 µg/20 µl | $120.00 | |
| D5030-1 | Lightning Conversion Reagent | 1.5 ml | $30.00 | |
| D5001-3 | M-Binding Buffer | 20 ml | $18.00 | |
| D5001-4 | M-Wash Buffer | 6 ml | $13.00 | |
| D5030-5 | L-Desulphonation Buffer | 10 ml | $20.00 | |
| D3004-4-1 | DNA Elution Buffer | 1 ml | $13.00 | |
| W1001-1 | DNase/RNase-Free Water | 1 ml | $12.00 | |
| C1004-50 | Zymo-Spin IC Columns | 50 Pack | $64.00 | |
| C1001-50 | Collection Tubes | 50 Pack | $18.00 | |
Description
Performance
Technical Specifications
| DNA Input | For optimal results, use 100 ng of high quality, intact genomic DNA as input. Protocol is compatible with inputs between 10 ng to 100 ng. DNA should be free of enzymatic inhibitors and RNA contamination. DNA can be resuspended in water, TE, or a low salt buffer. |
|---|---|
| Processing Time | ~ 4 hours |
| Required Equipment | Microcentrifuge, thermal cycler with heated lid, magnetic separator, multichannel pipette (suggested) |
| Required material not provided | Illumina DNA Prep, (M) Tagmentation Kit (Illumina, Cat. No. 20060059 or 20060060) |
| Sequencing Platform | Libraries are compatible with all Illumina sequencing platforms (MiSeq, MiniSeq, HiSeq, NextSeq, NovaSeq) |
Resources
Documents
FAQ
No. The Illumina DNA Prep Kit (Cat. No. 20018704 or 20018705) is required and must be purchased directly from Illumina.
No. The kit provides twelve (12) Unique Dual Index Primer Sets. If you require additional index primer sets, please contact our Technical Support for assistance (949) 679-1190 ext. 3 or tech@zymoresearch.com.
The size of the input genomic DNA is important for high-quality and high-yield library. DNA from FFPE samples are often highly degraded, so the quality of the library will be affected. Cell-free DNA are not compatible with the kit because the fragment size is too short for efficient tagmentation.
Zymo-Seq WGBS Library Kit is compatible with genomic DNA extracted from any species, whether it is mammalian, bacteria, or plants. It is best to use high-quality, intact genomic DNA that is RNA-free.
Libraries are suitable for any cycling number, but increased cycling numbers will require greater amount of adapter trimming. For most, 100 bp paired-end reads are suitable. The sequencing depth will be dependent on the genome size and coverage desired. Generally, aiming for 10x coverage per CpG site is recommended.
The number of reads (N) required for the desired coverage can be estimated by using the following equation: N = (Coverage)*(Diploid Genome Length)/Read Length. Bisulfite conversion reduces the complexity of the genome and have lower alignment rates compared to DNA-seq, so it is recommended to increase the number of reads by 25%. Therefore, for human WGBS at 10X coverage using 100 bp PE sequencing, at least 400 million reads are recommended.
Zymo-Seq WGBS kit is ideal for those working with higher inputs (10-100ng) of genomic DNA. It has minimal handling and clean-up steps, which makes it ideal for high-throughput library generation.
Pico Methyl-Seq would be a better option for those working with small DNA inputs (< 10ng) or highly fragmented DNA, such as cfDNA or FFPE DNA.
Citations
Need help? Contact Us