Successfully Added to Cart
Customers also bought...
-
 DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the...
DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the... -
 DNA/RNA Shield SafeCollect Swab Collection Kit (1 ml fill) (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA...
DNA/RNA Shield SafeCollect Swab Collection Kit (1 ml fill) (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA...
Highlights
- Quick recovery of ultra-pure DNA from agarose gels.
- Column design permits DNA elution at high concentrations into minimal volumes.
- Eluted DNA is well-suited for use in DNA ligation, sequencing, labeling, PCR, etc.
Original Manufacturer
100% satisfaction guaranteed, read Our Promise
Innovated in California, Made in the USA
Highlights
- Quick recovery of ultra-pure DNA from agarose gels.
- Column design permits DNA elution at high concentrations into minimal volumes.
- Eluted DNA is well-suited for use in DNA ligation, sequencing, labeling, PCR, etc.
Original Manufacturer
100% satisfaction guaranteed, read Our Promise
Innovated in California, Made in the USA
| Cat # | Name | Size | Price | |
|---|---|---|---|---|
| C1001-50 | Collection Tubes | 50 Pack | $18.00 | |
| C1004-50 | Zymo-Spin IC Columns | 50 Pack | $64.00 | |
| C1003-50 | Zymo-Spin I Columns | 50 Pack | $60.00 | |
| D3004-4-4 | DNA Elution Buffer | 4 ml | $12.00 | |
| D3004-4-1 | DNA Elution Buffer | 1 ml | $13.00 | |
| D4003-2-6 | DNA Wash Buffer (Concentrate) | 6 ml | $12.00 | |
| D4001-1-100 | ADB (Agarose Dissolving Buffer) | 100 ml | $76.00 | |
| D4001-1-50 | ADB (Agarose Dissolving Buffer) | 50 ml | $40.00 | |
| D4003-2-24 | DNA Wash Buffer (Concentrate) | 24 ml | $40.00 | |
Description
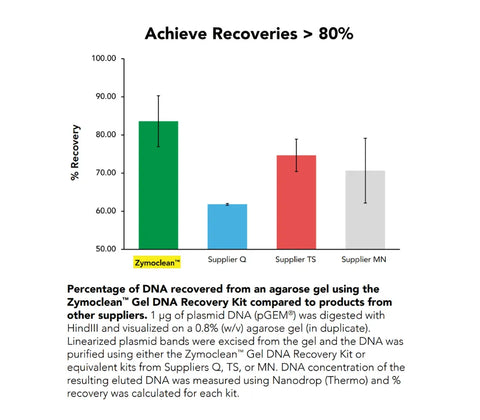
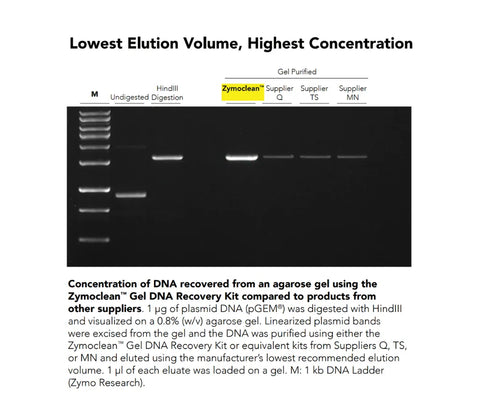
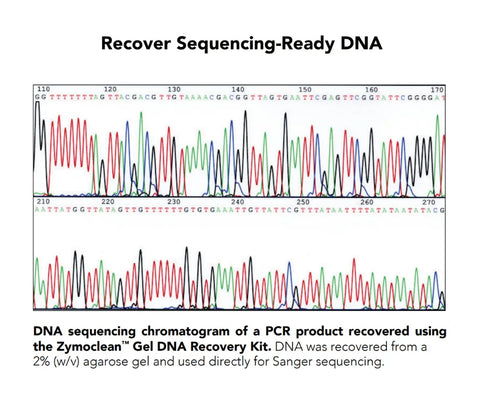
Performance
Technical Specifications
| Applicable For | Next Generation sequencing, ligation reactions, PCR, labeling, and restriction endonuclease digestions. |
|---|---|
| Elution Volume | ≥ 6 µl of DNA Elution Buffer |
| Equipment | Microcentrifuge |
| Purity | A260/A280 >1.8, A260/A230>1.8 |
| Sample Source | DNA excised from TAE/TBE-buffered agarose gels. |
| Size Range | 50 bp to 23 kb |
| Yield | Typically, up to 5 µg total DNA per column can be eluted with ≥6 µl water. For DNA 50 bp to 10 kb the recovery is 70-90%. For DNA 11 kb to 23 kb the recovery is 50-70% |
Resources
Documents
FAQ
The only difference is the cap. Everything else between the columns (max capacity, column matrix, etc.) is the same. Unsure which to use? It mainly comes down to preference. Some people like the capped columns for easy labeling or added security while others like the uncapped columns for faster sample processing.
Zymoclean has been validated for use with Zymo-Spin IIC and IIN columns. Other columns with higher binding capacity have not been validated by Zymo Research but have successfully been used by users.
Yes. Zymo-Spin I/IC Columns and DNA Wash Buffer are interchangeable.
Add 4 volumes of Agarose Dissolving Buffer for agarose gels >2% and ensure the gel slice has been fully dissolved.
No, high efficiency recovery is not guaranteed and the success of the purification can be variable.
If only isolating ssDNA, Zymo Research recommends the Zymoclean Gel RNA Recovery Kit. With some modification of the Zymoclean Gel DNA Recovery protocol, it is possible to recover ssDNA: After gel dissolving (step 3), add half a volume of isopropanol. Continue with the protocol.
Up to 400 mg of agarose gel.
Reviews
"The best advantage about this kit is that it is really quick to make it, and the total DNA recuperated have a good quality and it have a good concentration to follow the experiments."
Ingrid M.
"The kit was very easy to follow and yielded good results. It was more affordable than the Qiagen Gel Extraction kits, but worked just as efficiently. Solid product"
Thomas B.
University of Tennessee, Knoxville
"Protocols are very well written and Instructions easy to follow. Always had good results with your products."
Simin H.
University of California, Irvine
Citations
Need help? Contact Us