Successfully Added to Cart
Customers also bought...
-
 DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the...
DNA/RNA Shield (50 ml)Cat#: R1100-50DNA/RNA Shield reagent is a DNA and RNA stabilization solution for nucleic acids in any biological sample. This DNA and RNA stabilization solution preserves the... -
 DNA/RNA Shield SafeCollect Swab Collection Kit (1 ml fill) (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA...
DNA/RNA Shield SafeCollect Swab Collection Kit (1 ml fill) (1 collection kit)Cat#: R1160The DNA/RNA Shield SafeCollect Swab Collection Kit is a user-friendly collection kit for stabilizing the nucleic acid content of samples collected with a swab. DNA/RNA...
Highlights
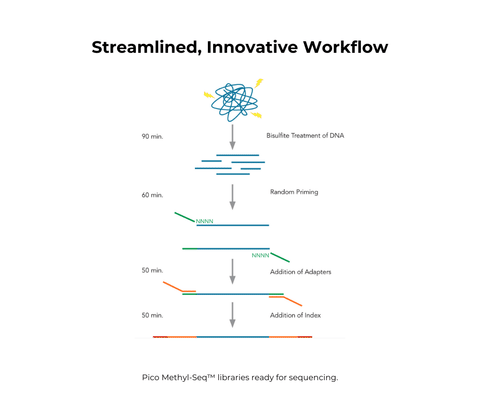
- All-inclusive kit for bisulfite conversion followed by Whole Genome Bisulfite Sequencing (WGBS) library preparation.
- Accommodates ultra-low DNA input and compatible with FFPE samples.
- Simple, ligation- and gel-free workflow can be completed in a few hours.
Original Manufacturer
100% satisfaction guaranteed, read Our Promise
Innovated in California, Made in the USA
Highlights
- All-inclusive kit for bisulfite conversion followed by Whole Genome Bisulfite Sequencing (WGBS) library preparation.
- Accommodates ultra-low DNA input and compatible with FFPE samples.
- Simple, ligation- and gel-free workflow can be completed in a few hours.
Original Manufacturer
100% satisfaction guaranteed, read Our Promise
Innovated in California, Made in the USA
| Cat # | Name | Size | Price | |
|---|---|---|---|---|
| D4003-2-6 | DNA Wash Buffer (Concentrate) | 6 ml | $12.00 | |
| D4003-1-25 | DNA Binding Buffer | 25 ml | $25.00 | |
| D5001-3 | M-Binding Buffer | 20 ml | $18.00 | |
| W1001-1 | DNase/RNase-Free Water | 1 ml | $12.00 | |
| D3004-4-1 | DNA Elution Buffer | 1 ml | $13.00 | |
| C1001-50 | Collection Tubes | 50 Pack | $18.00 | |
| C1004-50 | Zymo-Spin IC Columns | 50 Pack | $64.00 | |
| D5030-1 | Lightning Conversion Reagent | 1.5 ml | $30.00 | |
| D5030-5 | L-Desulphonation Buffer | 10 ml | $20.00 | |
| D5001-4 | M-Wash Buffer | 6 ml | $13.00 | |
Description
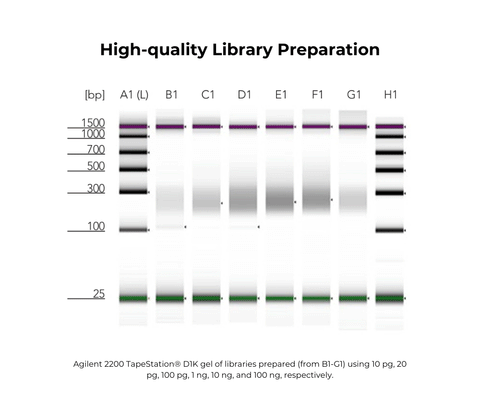
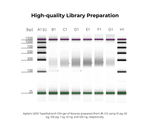
Performance
Technical Specifications
| Equipment | Microcentrifuge, thermocycler |
|---|---|
| Input | 10 pg - 100 ng |
| Sample Source | The protocol is designed for 10 pg – 100 ng genomic DNA input. DNA should be free of enzymatic inhibitors and can be suspended in water, TE, or a low salt buffer. DNA with low 260/280 or 260/230 ratios should be purified prior to processing using the Genomic DNA Clean & Concentrator (D4010). |
| Sequencing Compatibility | This kit produces libraries with Illumina TruSeq Adapters that can be sequenced on any of the Illumina sequencing platforms. |
Resources
Documents
FAQ
Yes, we have successfully prepared libraries from cfDNA using the Pico Methyl-Seq kit.
1. Ensure that there is diversity within the first couple of bases. Illumina recommends choosing barcodes that do not share the same base at the position. 2. Order new primers according to the “Can additional index primers be purchased for multiplexing?” section 3. Illumina recommends the following multiplexing strategy for 6 or fewer samples: Pool of 2 samples: • Index #6 GCCAAT • Index #12 CTTGTA Pool of 3 samples: • Index #4 TGACCA • Index #6 GCCAAT • Index #12 CTTGTA Pool of 6 samples: • Index #2 CGATGT • Index #4 TGACCA • Index #5 ACAGTG • Index #6 GCCAAT • Index #7 CAGATC • Index #12 CTTGTA
Yes, the index primer used in both library prep kits are the same. Both kits use the primer sequences off the Illumina TruSeq Small RNA Library Prep Kits.
We recommend aligning in the sequence in single read.
We typically sequence Pico Methyl-Seq libraries with 50 bp reads because trying to align reads that are longer than 50 bp result in lower alignment rates. If you still want to sequence the libraries at 100 bp or 150 bp reads, we suggest to trim the reads down to 50 bp to align, or sequentially 5-10 bp each time to see at what point the alignment goes up. Pico Methyl-Seq generated libraries are TruSeq libraries. The index barcodes are in the protocol and can be used in the sample sheet to upload onto the NextSeq for demultiplexing.
We quantify input DNA using Qubit or Nanodrop, but any method will do. As the range of input vs. cycles is wide, an estimation for input is sufficient. Based on 1 ng, we recommend running 10 cycles for amplification. The input should be RNA-free DNA.
We suggest setting the lid temperature not higher than 40°C for the PrepAmp reaction (see page 5, step 3 of section 2) as it might result in fragmentation.
Using a higher ratio of PrepAmp Primer to DNA input can result in primer dimers that get carried through to the final library. To help avoid banding pattern: - Try reducing the amount of PrepAmp Primer to 20 nM (for inputs of ~10-50 ng) or 10 nM (for inputs <10 ng). - Please evaluate the best ratio for your sample type and sample amount. Ratio dependent on type of input (gDNA, restriction-enzyme digested, cfDNA, etc.). Are libraries produced by the Pico Methyl-Seq library prep kit directional or non-directional? Libraries prepared with this kit are non-directional and as such, the original-top, original-bottom, and the complementary strands for each will be represented.
Yes, the EZ DNA Methylation – Lightning reagents can be substituted with the reagents from the EZ DNA Methylation – Direct kit (D5020).
The sequences of these primers are proprietary but are based off of the Illumina TruSeq adaptors. For trimming purposes, our bioinformatics team uses this sequence: GATCGGAAGAGC
Citations
Need help? Contact Us